|
Neighborhood Quality Standard (NQS) algorithm of (also see for more information).īased on your specifications on what you consider a valid SNP, the The SNP detection in CLC Genomics Workbench is based on the Single-nucleotide variations, CLC Genomics Workbench offers Manually checking all the conflicts of a contig to discover significant f) Discard reads below a certain length.ġ4) Single Nucleotide Polymorphism (SNP) detection - Instead of.d) Trim contamination from saved sequences.c) Trim contamination from vectors in UniVec database.Offers a number of ways to trim and filter out sequence reads prior to Sequence Number of reads Average coverage and Total number ofġ3) Trimming and filtering sequences - CLC Genomics Workbench The table includes the following information: Length of consensus.Generate a lot of contigs, and this option creates a table which makes itĮasier to get an overview of all the contigs. c) Table including all contigs: de novo assembly can potentially.b) List of non-assembled sequences: This will put all the reads thatĬould Not be assembled into a sequence list.a) Assembly report: This will generate a summary report.Output - CLC Genomics Workbench allows for three (3) types of output reporting: Full integration of the viewers included in the downstream analyses.ġ2) Quality reporting and statistics on raw data - Reporting of assembly (see below), making assemblies very fast.ġ1) Interactive and zoom-able viewing of genome assemblies, including sequencing reads, quality data, and reference sequences. exons.ġ0) Integration with CLC bio’s High Performance computing solutions ĩ) Masking of reference assembly based on annotations like e.g. Preparation of the sample for sequencing. One method is to tag the sequences with a unique identifier during the Throughput processes there is a need for being able to input severalĭifferent samples to the same sequencing run. Reads, so that the reads from one sample are assembled together.Ĩ) Support for Multiplex Sequencing by Tag - With many of the new high. There is often a data analysis challenge to separate the sequencing d) Paired end reads - duplications and inversions - CLC Genomics Workbench includes a number of graphical options of identifying genomic duplications and inversions when the sequencing produces paired-end reads.ħ) Multiplex Sequencing by Name - When you do batch sequencing ofĭifferent samples, you can use multiplexing techniques to run different.c) Paired end reads - insertions and deletions - CLC Genomics Workbench includes a number of graphical options of identifying genomic insertions and deletions when the sequencing produces paired-end reads.Much more advanced approaches to detecting genome rearrangements than single reads, and CLC Genomics Workbench therefore facilitates several ways of analyzing such paired end data. b) Paired-ends reads - graphical overview - Paired-ends data allows for.'coverage graph' along the contig by clicking the checkbox in the Side To assist in this interpretation, CLC Genomics Workbench displays a.Reads data, coverage is one of the main resources for interpretation. a) Single reads - coverage and conflicts - When you only have single.454 and IlluminaĤ) Reference assembly of genomes of any size.ĥ) Assembly of standard read data and support for assembly of pairedĮnd reads / mate pair reads of any sequencing technology.Ħ) Advanced graphical tools for the detection of large scale mutations Reads, and it supports Sanger, 454, Solexa, Helicos, and SOLiDģ) Reference assembly of mixed datasets (e.g. Second, all the readsĪre assembled using the contig sequence as reference.Ģ) Reference assembly - The reference assembly of CLC Genomics 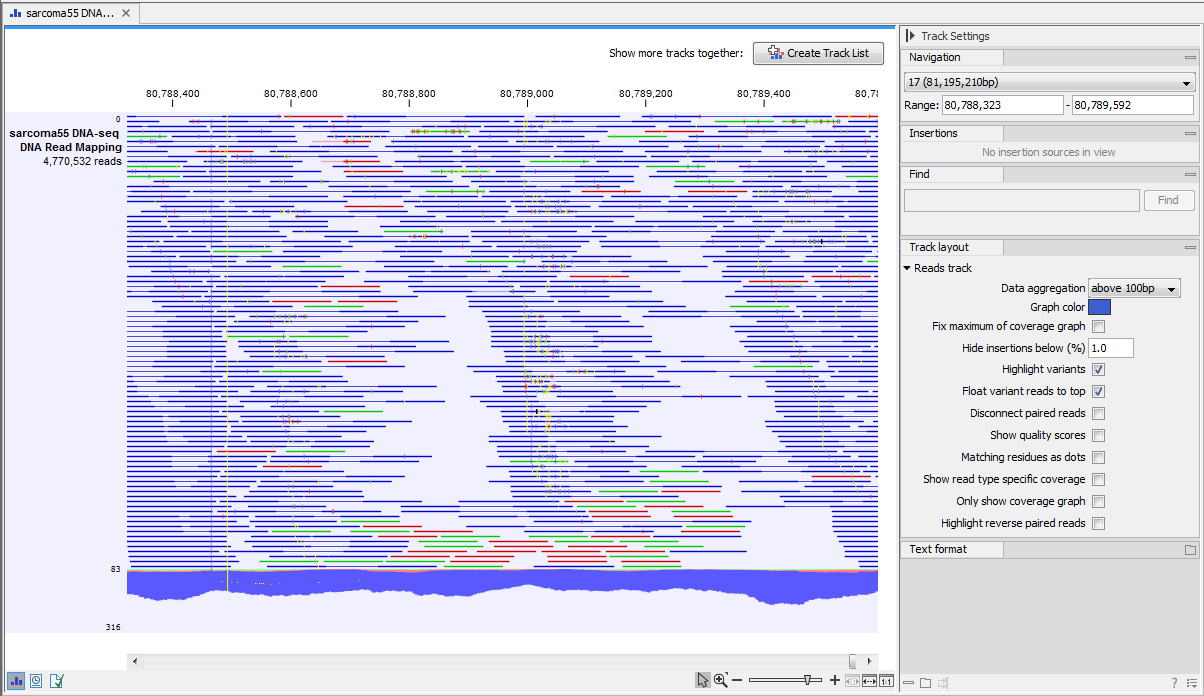
Sequences are created by aligning all the reads. The de novo assembly process has two stages: First, contig 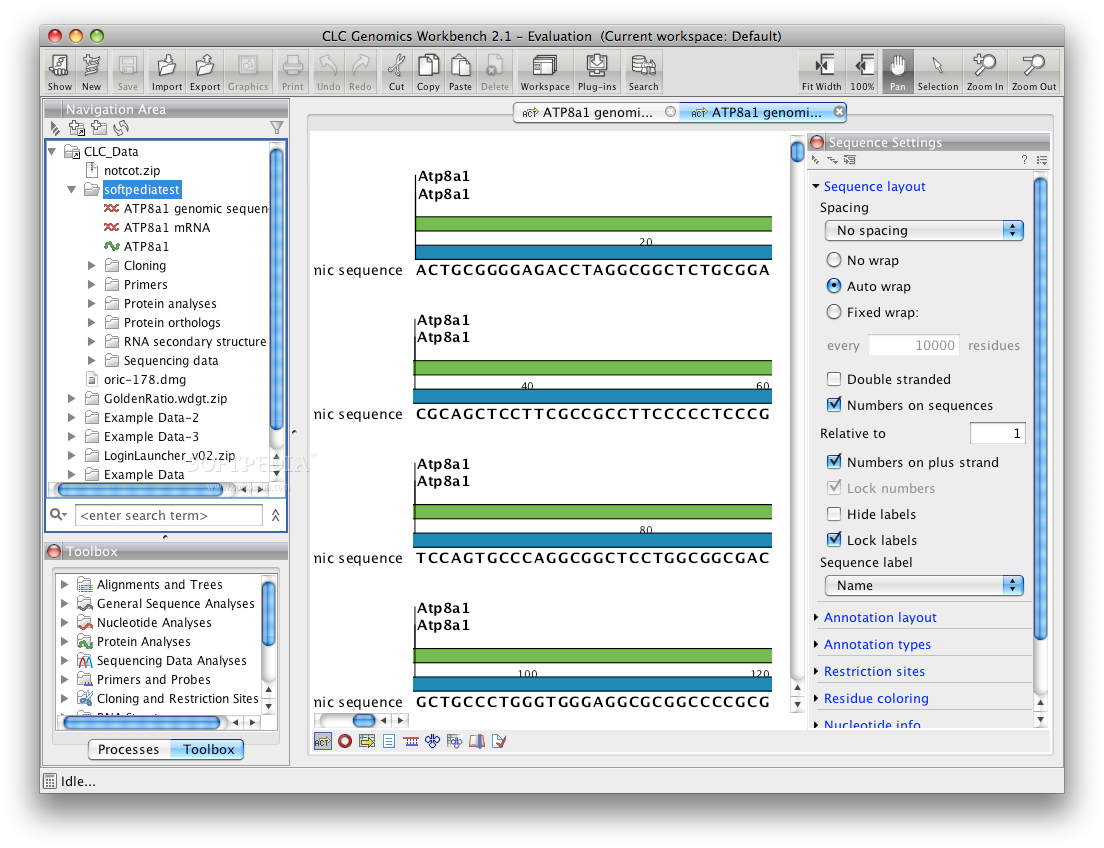
Reads, and it supports Sanger, 454, Illumina Genome Analyzer, Helicos, Workbench supports both short and long reads, it supports paired-ends Workbench' (see G6G Abstract Number 20096A) and the followingġ) De novo assembly - The de novo assembly of CLC Genomics Integrating with the rest of your typical NGS workflow.ĬLC Genomics Workbench includes all features of 'CLC Main It incorporates cutting-edge technology and algorithms, while also supporting and Category Cross-Omics>Next Generation Sequence Analysis/ToolsĪbstract CLC Genomics Workbench is a new solution for analyzing and visualizing Next Generation Sequencing (NGS) data.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
